Medizinische Epigenetik
Prof. Dr. Marc Timmers
Unsere Arbeitsgruppe untersucht die Transkriptionsregulation im Zusammenhang mit Chromatin-Modifikationen, die für humane Erkrankungen relevant sind. Im Besonderen konzentrieren wir uns auf die MLL-Komplexe, die für Histon-Methylierungen verantwortlich sind und die Aktivität von Gen-Enhancern und Gen-Promotoren regulieren. Gene, die für integrale Untereinheiten der MLL3- und MLL4-Komplexe kodieren, weisen die höchste Mutationsrate beim Blasenkrebs auf. Dieser Tumortyp ist durch ein starkes Geschlechterungleichgewicht gekennzeichnet, dessen mechanistische Grundlagen wir erforschen. Neben diesen molekularen Studien führen wir sowohl genetische als auch chemische Screens durch, um angreifbare Schwachstellen von Blasenkrebs-Zelllinien sowie primären 2D- und 3D-Kulturen aus Blasenkrebs zu bestimmen. Ziel ist die Entwicklung neuer Behandlungsoptionen für diese Erkrankung.
Die Menin-Untereinheit der MLL1/MLL2-Komplexe ist zu einem Schwerpunkt der Leukämieforschung geworden, da chemische Menin-MLL-Inhibitoren bei bestimmten Formen der AML bemerkenswerte Wirksamkeit zeigen und voraussichtlich bald von der FDA für die Behandlung zugelassen werden.
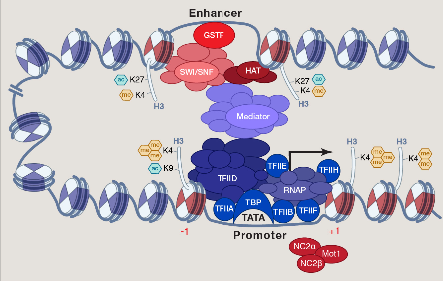
Alle zellulären und epigenetischen Signalwege führen letztlich zur Bindung der RNA-Polymerase II an spezifische Gen-Promotoren. Der basale Transkriptionsfaktor TFIID spielt dabei eine koordinierende Rolle bei der Regulierung der Transkriptionsinitiation. Wir untersuchen die Chromatin-Aspekte der dynamischen Regulation der TFIID-Aktivität.
Generell nutzen wir für unsere Studien zur Gen-Transkription und Chromatinregulation Kombinationen aus Zellbiologie, Molekularbiologie, Proteomik, Genomik, automatisierter Mikroskopie, Bioinformatik sowie chemischen und genetischen Bibliotheken mit Fokus auf Epigenetik.
Zukünftige Projekte und Ziele
Wir wollen epigenetische Mechanismen und therapeutische Schwachstellen beim Blasenkrebs identifizieren. Mithilfe von KDM6A/UTX und UTY als Modell-Gene möchten wir die Geschlechterdifferenz bei Blasenkrebs verstehen. Die MLL3- und MLL4-Histon-H3K4-Methylase-Komplexe stehen im Zentrum unserer mechanistischen Untersuchungen. Besonderes Interesse gilt der epigenetischen Signalübertragung von MLL-Komplexen zum basalen Transkriptionsfaktor TFIID und dessen Regulation.

Prof. Dr. Marc Timmers
Professur – Partnerstandort Freiburg
Universitätsklinikum Freiburg – Abteilung für Urologie
Zentrale Klinische Forschung (ZKF)
Medizinische Epigenetik / Medical Epigenetics