Translational Oncology (Neurooncology)
Prof. Dr. Björn Scheffler
Cancer adapts. Under therapeutic pressure, it evolves, reshapes its microenvironment, and evades immune control. Our goal is to anticipate these adaptive dynamics and intercept them before therapeutic resistance becomes established.
Embedded within a strong clinical framework at the West German Cancer Center Essen (WTZ), we integrate direct investigation of human disease - through advanced molecular imaging and analysis of primary patient samples - with patient-derived stem cell platforms and orthotopic models.
Within this translational continuum, we track disease dynamics in patients, dissect oncogenic mechanisms, and translate biological insight into biomarkers and early therapeutic strategies.
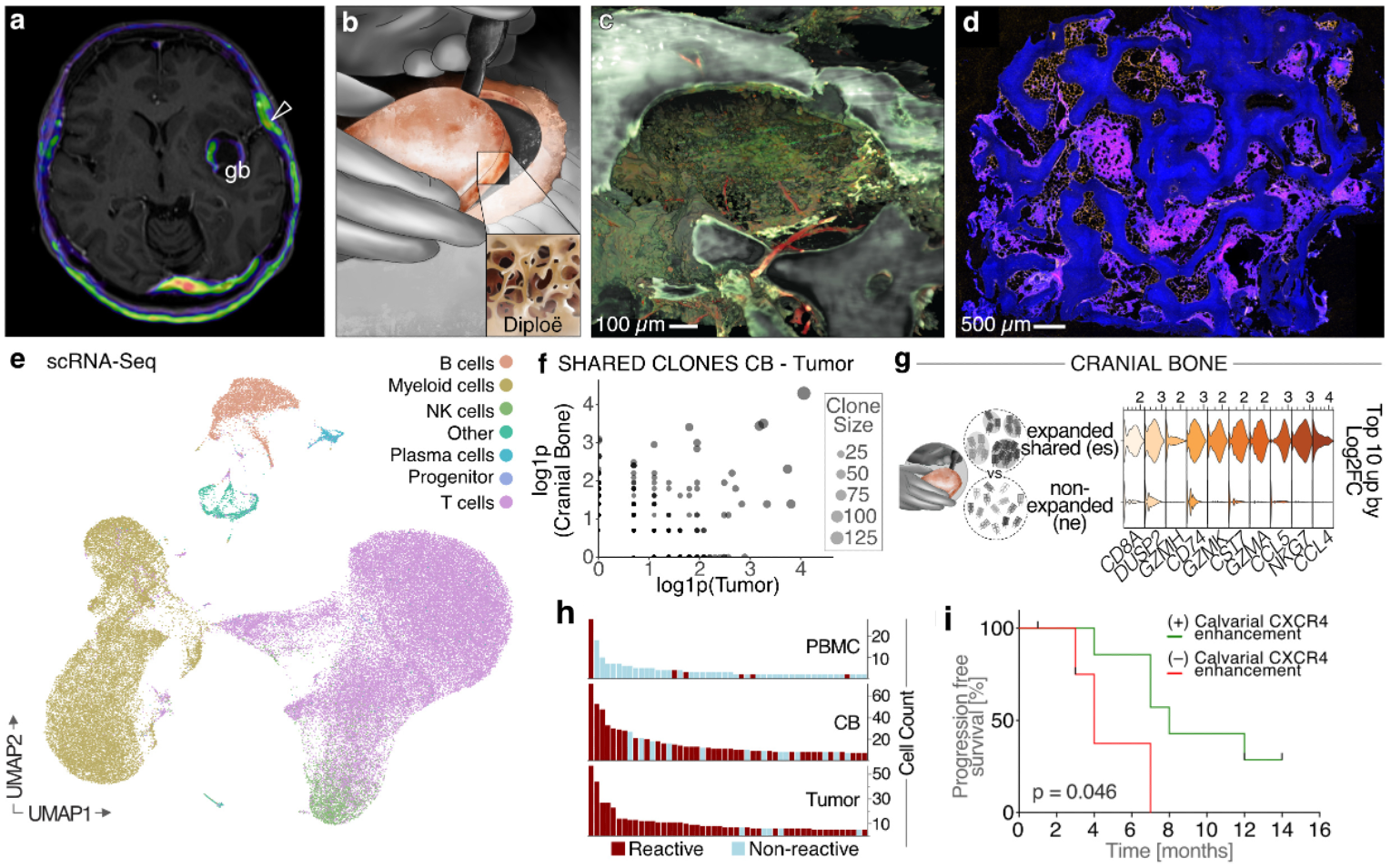
A central focus of our current work is the immune landscape at the brain’s borders. Our research has contributed to the recognition that brain tumors and the adjacent cranial bone-meningeal compartments constitute an active hematopoietic and immune niche directly interfacing with the CNS. Far from being a passive boundary, this cranial immune compartment appears to shape tumor evolution, immune surveillance, and therapeutic responsiveness.
By defining and mechanistically interrogating this tumor-cranial niche axis, we aim to enable refined biomarker platforms, localized immune modulation, and next-generation strategies for CNS-directed therapy.
More information can be accessed on this webpage.

Prof. Dr. Björn Scheffler
DKFZ Division of Translational Oncology / Neurooncology
West German Cancer Center (WTZ) University Hospital Essen
Translationale Onkologie (Neuroonkologie) / Translational Oncology (Neurooncology)